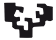Int. J. Dev. Biol. 66: 155 - 161 (2022)
Special Issue: Developmental Biology in Greece
Zebrafish research in Greece: swimming against the current
Review | Published: 20 September 2021
Abstract
The zebrafish is a vertebrate model extensively used in Developmental Biology and Human Disease modeling, as it shares high genetic and physiological similarities with humans. It has become the second most popular animal model, after mice, with several advantages over the latter: zebrafish are easily housed and cared for; the cost of installing and maintaining a zebrafish facility is significantly lower than for mice; and they reproduce often and develop quickly. Using zebrafish complies with the 3Rs principles of laboratory animal use. Zebrafish embryos develop externally and are transparent, allowing for in vivo non-invasive imaging. There are many transgenic and mutant lines available that mimic most human diseases, including reporter lines for most signaling pathways. There are also several reverse genetic tools to functionally verify genes or variants of unknown significance, identified in Genome-Wide Association Studies (GWAS) or using Next Generation Sequencing (NGS) approaches. In addition, the model emerges as an invaluable whole animal platform for various stages of drug discovery efforts by exploring the possibility of creating high-throughput phenotypic-driven screens. These include phenotypic screenings, determinations of general and/or specific toxicity (cardiac, renal, hepatotoxicity etc.), and mechanism of action studies. Finally, zebrafish are able to retain their capacity to regenerate most organs during their entire life span, making them a well-established model for the study of organ regeneration. The European Zebrafish Society consists of more than 180 research labs throughout Europe. In Greece however, zebrafish use remains rather limited. Here I present here a brief historical overview of zebrafish research in Greece.
Keywords
zebrafish, Human Disease models, toxicology, precision medicine, regeneration
Zebrafish in Developmental Biology
The extraordinary possibility to observe development non-invasively always amazes somebody who first looks through a stereoscope at a zebrafish embryo. The advances made in microscopy and, most importantly, in Spinning Disc Microscopy (spim) in combination with transgenic lines were pivotal in enabling the detection, imaging and reconstruction of development and organogenesis at cellular resolution (Huisken et al., 2004; Keller et al., 2008). Advances in next-generation sequencing (NGS) techniques made it possible to reconstruct a transcriptomics-based single-cell lineage of development and organogenesis (Wagner et al., 2018; Farrell et al., 2018). Looking at live embryos and, more specifically, their beating hearts, was how I got hooked on zebrafish research. After my post-doctoral training at the University of California San Francisco (UCSF), where I participated in a forward genetic screen (Beis et al., 2005), I decided to move back to Greece and set up my own zebrafish facility at the Biomedical Research Foundation of the Academy of Athens (BRFAA) in 2005 (Fig. 1). At that time, with the Stainier lab team, I had just witnessed the relocation and reinstallation of a large zebrafish facility from the Parnassus campus to Mission Bay, so moving some fish across the ocean did not seem such a challenge.
During the first years after establishing my lab in BRFAA, we primarily focused on cellular and developmental biomedical approaches to cardiovascular development and disease. To my knowledge, I was the first person to introduce zebrafish biomedical research in Greece, and I still maintain the largest housing facility. The ease of studying cardiovascular development and angiogenesis in vivo was soon appreciated, and in 2006, in an initial collaborative paper with Greek research teams, we showed that activation of soluble Guanylyl Cyclase (sGC) promotes angiogenesis. (Pyriochou et al., 2006). Since then, we have extensively used the Tg(kdrl::eGFP) line to screen for angiogenesis inhibitors in several projects. We used it to select molecules with the maximal activity of VEGFR2 inhibition (Papakyriakou et al., 2014) and peptide-conjugated molecules synthesized to target cancer cells (Argyros et al., 2018).
Zebrafish Models of Cardiovascular Development and Disease
The zebrafish heart provides a powerful in vivo system to study cardiac development and regeneration at the cellular level (Bournele and Beis 2016; Giardoglou and Beis 2019). We used cardiac valve development to dissect the role of intracardiac flow dynamics in mutants with unlooped hearts (southpaw) and non-contracting atrium (myh6, weak atrium) (Kalogirou et al., 2014). The fascinating phenotype of myh6 adults that survive with a single functioning chamber allowed us to study a model of adult ventricular remodeling. We showed that the increased ventricular size is due to a hyperplastic response, besides exhibiting a transcriptional program that in mammals is mostly associated with hypertrophy (Sarantis et al., 2019). Currently, we are also studying a model of total aortic stenosis that carries a mutation in protein kinase d2 (prkd2) (Giardoglou et al., 2021). prkd2 mutants initiate blood circulation but fail to maintain it, allowing the study of valve development in the absence of proper intracardiac blood flow pattern, a phenotype that would be instantly lethal in mammalian systems (Fig. 2). Simultaneously, in a collaborative study with research groups at the University of Crete, we showed that early developmental exposure to diverse temperatures could be associated with a different cardiac morphology. Increasing the temperature during development caused ventricular roundness and decreased exercise performance (Dimitriadi et al., 2018) of adult fish. Our collaborators had already shown (Georga et al., 2010) that early and relatively limited periods of exposure to different temperatures during development could affect the body shape and several morphological characteristics, including caudal fin segmentation pattern (Christou et al., 2018), which are also relevant to aquaculture. Recently, they also set up a micro-CT scanning-based protocol and 3D morphometrics to quantify in detail the changes in heart morphology (Dimitriadi et al., 2021).
Fig. 2. prkd2 mutants exhibit a complete outflow tract stenosis.
Confocal image of an embryonic heart from prkd2 (protein kinase D2) mutant zebrafish depicting the cardiac endothelium with Tg(kdrl::eGFP) and circulating blood cells with Tg(gata1::DsRed). Panels are pseudocolored, inspired by Andy Warhol’s art movement. prkd2 embryos exhibit the extreme phenotype of outflow stenosis (arrow) and total retrograde blood flow from the ventricle to the atrium and no blood circulation by 72 hours post fertilization (hpf). They survive however, with a beating heart, for up to 120 hpf.
Zebrafish Models of Neurobiology
In the early days of zebrafish research in Greece, there was also interest in and work focused on adult neurogenesis. In a series of studies, it was shown that there is sexual dimorphism in cerebellum cell proliferation (Ampatzis and Dermon 2007). BrdU incorporation in specific brain areas, as well as the immunohistochemistry of several receptors in distinct brain regions, enabled researchers to unravel the specific distribution of beta (2)-adrenergic receptors. They were subsequently able to correlate this regional distribution with swimming behavior, anxiety-like behavior, novel tank exploration and habituation of adult zebrafish, as well as sexual dimorphisms (Ampatzis et al., 2008; Ampatzis and Dermon 2010; Ampatzis et al., 2012; Ampatzis and Dermon 2016).
A significant contribution of Greek zebrafish research in zebrafish behavior and husbandry was the introduction of a very sensitive protocol for measuring a vital stress marker (cortisol measurement) (Pavlidis et al., 2013; Samaras and Pavlidis 2020). The group proceeded to determine the molecular mechanisms underlying the stress response, including crf (corticotropin-releasing factor) and bdnf (brain-derived neurotrophic factor) transcriptional dynamics (Pavlidis et al., 2015). These analyses, together with morphological malformations that were caused in a model of exercise-induced lordosis in juvenile zebrafish (Printzi et al., 2021), also brought zebrafish research closer to aquaculture-species physiology. Research groups at the University of Crete, together with colleagues at the Hellenic Centre for Marine Research, studied the physiology of infections from a bacterial family related to aquaculture: Edwardsiella. Such outbreaks account for severe biomass losses and financial damage (Katharios et al., 2019), making zebrafish research more relevant for aquaculture, an important area for the Greek economy.
Zebrafish Models of Regeneration
It is becoming increasingly evident that the reactivation of developmental pathways is necessary to drive regeneration. Although adult mammalian hearts have limited regenerative capacity, newborn mice have the ability to regenerate. Consequently, the hypothesis that re-activating or suppressing specific signaling pathways after injury could stimulate endogenous regenerative potential in mammals continues to be a thought-provoking notion. My group studied cardiac valve regeneration in zebrafish, using an inducible tissue-specific genetic ablation model. We expressed the Nitroreductase gene (UAS: Nitroreductase) in a valve-specific pattern, and by adding metronidazole at various time points, we induced valvular damage. We established that zebrafish cardiac valves can regenerate, and isolated regenerating hearts to identify genes that are differentially regulated upon valvular damage using RNASeq. We characterized Notch and TGFβ signaling as indispensable during valve regeneration (Kefalos et al., 2019; Bensimon-Brito et al., 2020).
The zebrafish has also proven to be an excellent model because of its amazing regenerative potential to functionally heal a spinal cord injury (SCI). In collaboration with several groups in Germany, we have analysed how perivascular cells, and the specific ECM components they secrete, promote regeneration in zebrafish, while similar cells create a non-permissive environment that leads to scar formation in mammals following SCI (Tsata et al., 2021).
Zebrafish in Translational Research and Precision Medicine
The high degree of genetic homology between zebrafish and humans (it is estimated that more than 70% of human disease-causing genes have a clear orthologue in zebrafish), has led to the development of several zebrafish human disease models. Furthermore, the ability to perform high-throughput, phenotypic-driven screens allows for the identification of potentially valuable preclinical molecules that could lead to drug development. A paradigm study for this approach is the generation of a zebrafish transgenic line carrying the 2057del2 mutation in the plakoglobin gene (Asimaki et al., 2014) that causes the Naxos disease (Protonotarios et al., 1986). Following the characterization of the phenotype, the authors conducted a high-throughput screen and identified a suppressor of cardiac mortality in fish. They then continued and verified these results in rat cardiomyocytes carrying the plakoglobin mutation (Asimaki et al., 2014).
My group was able to study a human adult-onset peripheral neuropathy: Charcot-Marie-Tooth (CMT), CMT2D caused by mutations in the Glycyl t-RNA synthetase (GARS). We identified and characterized a mutant line in the zebrafish gars where a Threonine to Lysine mutation in the dimerization domain inhibited protein dimerization and function. The embryos exhibited progressive loss of neuromuscular junctions (Malissovas et al., 2016). Using human characterized pathogenic alleles (all in conserved amino acids), we were able to show that dimerization between a mutant monomer and the wild type is required for the autosomal dominant disease, and indicate that a dominant-negative effect may be a potential mechanism of GARS-mediated CMT disease. This also provided us with a model in which new variants of unknown significance in the GARS gene could be tested in vivo, and where one could also search for suppressors of the phenotype. This approach is currently being exploited by several human genetics groups studying variants of unknown function or novel genes in human development. For example, mutations in DYNC1I2 (Dynein Cytoplasmic 1 Intermediate Chain 2), encoding a component of the cytoplasmic dynein 1 complex, was first identified as a putative candidate gene in a Pakistani family with microcephaly and intellectual disabilities, and was shown in zebrafish to cause reduced head size (Ansar et al., 2019). In addition, mutations in the DDX3X (DEAD-Box Helicase 3 X-Linked) were identified that putatively cause intellectual disabilities in two male siblings. DDX3X is necessary for canonical Wnt signaling, and therefore there is a functional assay to evaluate DDX3X variants. It is based on their ability, when coinjected with Wnt, to interact and cause ventralization of zebrafish embryos. This observation allowed for the functional verification and characterization of a novel variant of unknown significance as a hypomorph, providing in vivo evidence of this rare genetic disease (Kellaris et al., 2018). The CRISPR/Cas9 technology has revolutionized reverse genetic approaches and an increasing amount of knock-out and knock-in zebrafish lines are being generated to functional study genes of interest. We have studied a 3-mercaptopyruvate sulfurtransferase (3-mst) mutant where we detected elevated levels of reactive oxygen species (Katsouda et al., 2020) and a protein tyrosine phosphatase receptor-ζ1 ptprz −/− knockout, which exhibited misregulated expression of developmental cardiac markers, bradycardia, and defective heart morphogenesis characterized by enlarged ventricles and defected contractility (Katraki-Pavlou et al., 2022).
Besides human genetic analyses using zebrafish, my group has also been involved in several medium to high-throughput phenotypic screens to identify novel bioactive compounds derived from various sources, including natural compound extracts. We have shown that crocins can regulate sugar levels in vivo using whole zebrafish embryo sugar measurements and the pancreas-specific transgenic line Tg(ins::DsRed) (Kakouri et al., 2020). Mastic essential oil, a highly valued product unique to Greece, has also been studied in vivo using zebrafish. A group at the University of Ioannina analyzed the toxicological profile and immune response transcriptomic profile following addition of mastic essential oil to zebrafish embryo water, as well as its developmental effect on lateral line differentiation (Serifi et al., 2019). My lab has been systematically screening for inhibitors of melanocyte differentiation. During this process, which initially cannot differentiate between a melanin synthesis inhibitor and a melanocyte differentiation inhibitor, we identified and characterized two natural extracts from Morus alba and Craetegus pycnoloba. We showed that both work through melanin synthesis inhibition with potential usage in cosmetics, or as food supplements in aquaculture (Chaita et al., 2017; Agalou et al., 2018). We continue to search for melanocyte differentiation inhibitors with potential application in the treatment of metastatic melanoma.
Numerous zebrafish cancer models have been developed during the last ten years. The high degree of homology between humans and zebrafish allows the creation of transgenic zebrafish lines introducing human oncogenes under a tissue-specific promoter. Zebrafish cancer models, and especially the emerging powerful models of cancer cell xenotransplantations, can also be used to screen for cancer biosensors. We have tested in vivo such a molecule, which reacts with biothiols in cancer cells, emits near-infrared fluorescence and therefore could be used as a non-invasive cancer probe (Diamantis et al., 2021). Currently, these are well-established protocols in several labs, and the metastatic behavior of cancer lines as well as their angiogenic activity and immune-triggering responses can be followed in vivo. There is an excellent correlation between the zebrafish results with mammalian models and their in vivo response to treatment with existing chemotherapies. The main advantage of zebrafish is the non-invasive in vivo imaging of cancer progression, as well as the limited time needed to monitor these responses, which in most cases ranges from 4-10 days, compared to several weeks in mouse models. Zebrafish patient-derived xenografts (zPDX) could distinguish functional tumor behaviors in just 4 days and showed differential sensitivity to colorectal cancer therapy (Fior et al., 2017). Over the past twelve months, and using funding from the OpenScreenGR infrastructure grant from which the Beis lab benefits, we established the xenotransplantation protocol. We have successfully injected and tested the metastatic potential of several human cancer cell lines, including lung cancer and melanoma. We labeled cancer cells with the lipophilic dye DI and injected 200–10.000 cells depending on the line. In addition, we performed a pilot screen using the lung cancer cell line (A549) and the reference drug gemcitabine, showing inhibition of the engraftment, proliferation and metastasis of cancer cells following xenotransplantation in vivo (Fig. 3). It would significantly benefit the Greek research community to establish a protocol for zebrafish patient-derived xenografts (also known as zebrafish avatars) (Costa et al., 2020) that could permit patient-specific optimization of cancer therapy.
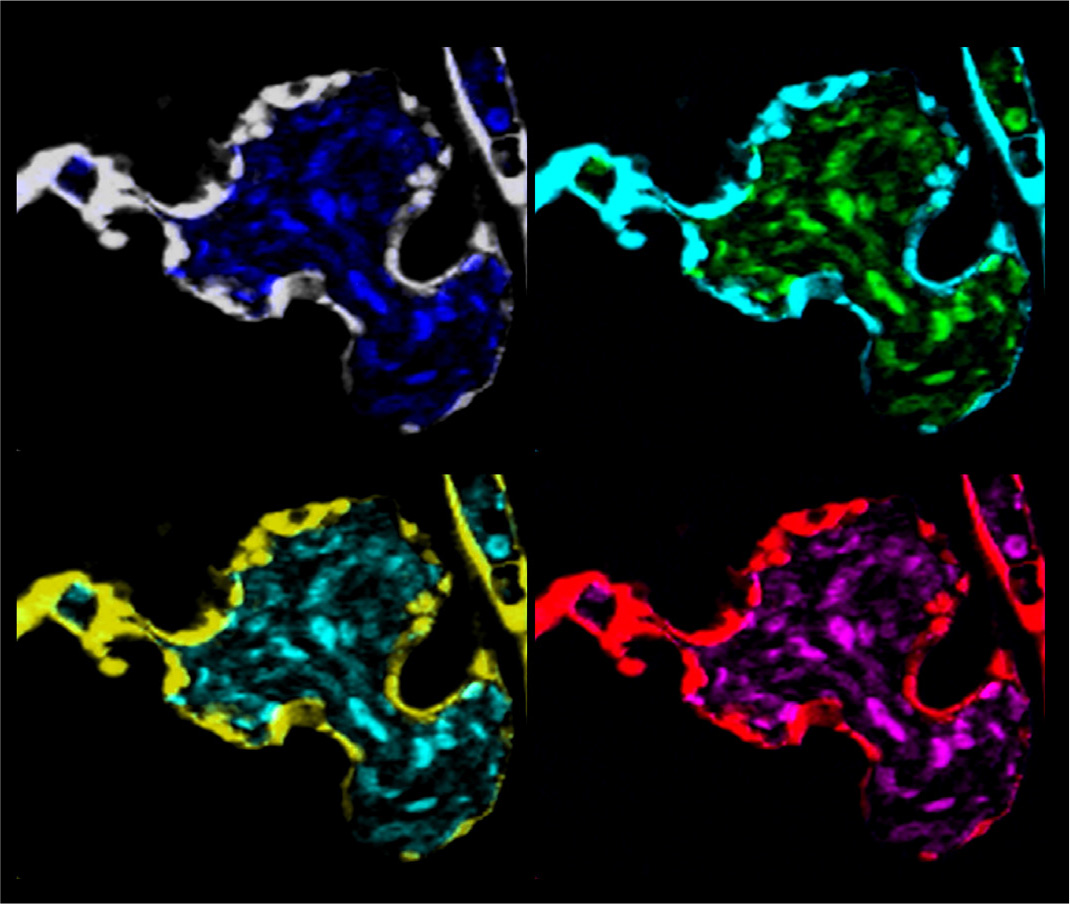
Fig. 3. Gemcitabine inhibits the growth of A549 cancer cells in vivo.
We labeled the lung cancer cell line (A549) with the lipophilic dye DI and injected ~200cells per embryo at two days post fertilization (dpf). Using the reference drug gemcitabine at 10μΜ, we showed inhibition of the engraftment, proliferation and metastasis of cancer cells in vivo within five days post injection (dpi). This approach can be used for any cancer cell line or zebrafish Patient-Derived Xenografts (zPDX).
Zebrafish in toxicology
In phenotype-driven screens, a critical component for the characterization of extracts is estimation of their toxicity, in order to unravel and understand the actual underlying phenotype. There is an OECD-approved fish embryo lethality test that is used to determine lethal concentration (LC) parameters such as LC50 (OECD 236). Several Greek labs have used the zebrafish model to study the toxicological properties of various compounds. My group has characterized the in vivo effects during the development of the emerging pollutants, Benzotriazoles (Damalas et al., 2018).
Several other groups have used zebrafish embryo screens as a toxicology model. At the University of Crete, they studied the toxicological properties and effects of waterborne Cadmium on adult fish, unraveling an unexpected non-linear response to Cd (Renieri et al., 2017), and at the Benaki Phytopathological Institute in Athens, they are integrating the zebrafish embryo test in their standard developmental toxicology analyses. They have shown that triadimefon (a widely used triazole fungicide) causes developmental defects (Zoupa and Machera 2017). The zebrafish embryo test can also be easily used to study the additive effects of different chemicals used in mixtures. This can be tested in a series of doses to check for morphological defects or lethality as an end-point (Zoupa et al., 2020).
Groups at the University of Ioannina tested the cardiotoxic effects of the anticancer drug adriamycin (Orfanidou et al., 2013) and evaluated the toxicity of two molecules with anticancer properties: N-(p-coumaroyl) serotonin (CS) (Lazari et al., 2017); and deglucohellebrin (DGH), which is extracted from the roots of Helleborus (Vartholomatos et al., 2020). They also studied the effects of cannabidol (a degradation product of the cannabis metabolite Δ9-tetrahydrocannabinol) during development as well as in survival and the metabolism of zebrafish embryos (Chousidis et al., 2020a). A challenge with novel compounds used in industry is the evaluation of their toxicity. The impact of carbon nanodots (CNDs) and carbon quantum dots (CQDs) was estimated using the metabolic profile and locomotor behavior of zebrafish embryos (Chousidis et al., 2020b; Chatzimitakos et al., 2021). In parallel, they also analyzed the transcriptional and behavioral impact on zebrafish embryos of a potent toxin produced by cyanobacteria: microcystin-LR (MCLR) (Papadimitriou et al., 2012; Tzima et al., 2017). A study with potential implications for local environmental planning and monitoring of the Ioannina lake.
In a collaborative effort between laboratories at the Aristotle University of Thessaloniki and the University of Crete, researchers also utilized zebrafish as part of their toxicology analyses. They characterized at high resolution the physiological response of several species to either titanium dioxide nanoparticles (TiO2-NPs) (Bobori et al., 2020) or magnetite nanoparticles (Kaloyianni et al., 2020). They were able to find specific biomarkers for each pollutant that were shared in several species. These biomarkers could be used to systematically monitor water pollution.
Zebrafish as an in vivo model for Biochemistry and Cell Biology
Since the early days of zebrafish research, it has become obvious that this will be a powerful model for Cell Biology and Biochemistry (Beis and Stainier, 2006). The advances in BioImaging and the transgenic lines generated over the years have provided all the necessary tools to exploit this potential to the fullest. Soon after the establishment of my lab in BRFAA, we were able to follow the phenotype of a G-protein coupled receptor regulating protein. Knockdown of regulator of G protein signaling 4 (RGS4), a G i/o GTPase-activating protein (GAP), allowed us to show its involvement in β-cell migration and islet aggregation during the development of pancreas (Serafimidis et al., 2011).
The generation of zebrafish transgenic lines with artificial promoters allowed imaging at cellular resolution of the activation of important signaling pathways. We participated in the developmental dissection of a widely used wnt signaling reporter line Tg(7xTCFXla.Siam:nlsmCherry) (Moro et al., 2012). We have also utilized the Notch reporter lines Tg(Tp1:venus-PEST) and the Tg(Tp1:h2B:mCherry) in several experiments, including valve regeneration (Kefalos et al., 2019). These lines show different kinetics of activation and signal maintenance due to the different fluorescent proteins they express. The destabilized venus-PEST is more dynamic, while mCherry can be used to visualize activation during longer periods. We also used the Hedgehog reporter line Tg (12xGliBS: mCherry-NLS), together with researchers at the University of Ioannina, to show that the oncoprotein (protein phosphatase 2A inhibitor 2) SET/I2PP2A inhibits Gli1 transcription (Serifi et al., 2021). Our collaborators at Ioannina have used zebrafish to functionally verify in vivo and support Biochemical data regarding several signaling pathways. They studied the effect of Prothymosin alpha (ProTα) in cell proliferation and apoptosis during zebrafish embryonic development (Emmanouilidou et al., 2013). They also studied the role of the oncoprotein SET/I2PP2A during development of the zebrafish sensory system. They showed differential expression patterns following knockdown of the set genes in several organs, including the eye retina, the ear and the lateral line (Serifi et al., 2016).
Outlook for zebrafish research in Greece
The International or European Zebrafish meetings are the most diverse meetings I attend, reflecting the wide-ranging potential for using zebrafish in research. I have attempted to highlight some of this potential. There are now several labs working with zebrafish, and the ever-increasing number of requests I receive for new collaborations demonstrates that more and more research groups are interested in embarking on zebrafish-based research. Once funding in Greece (especially for infrastructures) becomes regular it, will be crucial to establish a Greek Zebrafish Resource Center, with an open access policy to the Greek research community. This would be an essential addition to Greek Research Infrastructure for several reasons: a centralized zebrafish facility will be cost-effective, efficient and will help Greek research groups in numerous and very versatile projects; a centralized screening facility for Bioactive compounds, could boost zebrafish research and preclinical drug development programs in Greece; it would also be an ideal place to educate a new generation of Developmental Biologists in Greece. The zebrafish is particularly suited to hands-on laboratory research. Considering the versatility of small, easy to set-up experiments for undergraduates, it should also be integrated into the curriculum of every Biology or Medical department.
Acknowledgements
I would like to acknowledge Panayotis Karayannakos and Nikolaos Kostomitsopoulos for their help in setting up the zebrafish facility in 2005, and Katia Karalis for coordinating the TransMed REGPOT Eugrant (2008-2012) that allowed the maintenance and expansion of my facility. I would also like to thank Adamantia Agalou and Panagiota Giardoglou for the figures in this manuscript, as well as the past and present members of the Beis lab, undergraduate students, PhD candidates and post-doctoral fellows for taking such good care of the fish during the last fifteen years. Finally, a big thank you to the Fondation Santé for twice awarding me a Research Grant at very crucial moments in the history of my lab.
Abbreviations
bdnf, brain-derived neurotrophic factor ; BrdU, Bromodeoxyuridine ; BRFAA, Biomedical Research Foundation of the Academy of Athens ; crf, corticotropin-releasing factor) ; DDX3X, DEAD-Box Helicase 3 X-Linked ; DYNC1I2, Dynein Cytoplasmic 1 Intermediate Chain 2 ; GWAS, Genome-Wide Association Studies ; hpf, hours post fertilization ; kdrl, kinase insert domain receptor like ; myh6, myosin heavy chain 6 ; NGS, next-generation sequencing ; prkd2, protein kinase d2 ; ProTΑ, Prothymosin alpha ; SET/I2PP2A, protein phosphatase 2A inhibitor 2 ; sGC, soluble Guanylyl Cyclase ; spim, Spinning Disc Microscopy ; TGFΒ, transforming growth factor beta ; VEGFR2, vascular endothelial growth factor receptor-2 ; UAS, upstream activated sequence ; zPDX, Zebrafish patient-derived xenografts ;References
Agalou A., Thrapsianiotis M., Angelis A., Papakyriakou A., Skaltsounis A.L., Aligiannis N., Beis D. (2018). Identification of Novel Melanin Synthesis Inhibitors From Crataegus pycnoloba Using an in Vivo Zebrafish Phenotypic Assay. Front. Pharmacol. 9: 265.
Ampatzis K., Dermon C. R. (2007). Sex differences in adult cell proliferation within the zebrafish (Danio rerio) cerebellum. Eur J Neurosci 25: 1030-1040.
Ampatzis K., Dermon C. R. (2010). Regional distribution and cellular localization of β 2 -adrenoceptors in the adult zebrafish brain ( Danio rerio ) . J. Comp. Neurol. 518: 1418-1441.
Ampatzis K., Dermon C. R. (2016). Sexual dimorphisms in swimming behavior, cerebral metabolic activity and adrenoceptors in adult zebrafish (Danio rerio). Behav Brain Res. 312: 385-393.
Ampatzis K., Kentouri M., Dermon C. R. (2008). Neuronal and glial localization of α2A-adrenoceptors in the adult zebrafish (Danio rerio) brain. J. Comp. Neurol. 508: 72-93.
Ampatzis K., Makantasi P., Dermon C.R. (2012). Cell proliferation pattern in adult zebrafish forebrain is sexually dimorphic. Neuroscience 226: 367-381.
Ansar M., Ullah F., Paracha S. A., Adams D. J., Lai A., Pais L., Iwaszkiewicz J., Millan F., Sarwar M. T., Agha Z., Shah S. F., Qaisar A. A., Falconnet E., Zoete V., Ranza E., Makrythanasis P., Santoni F. A., Ahmed J., Katsanis N., Walsh C., Davis E. E., Antonarakis S. E. (2019). Bi-allelic Variants in DYNC1I2 Cause Syndromic Microcephaly with Intellectual Disability, Cerebral Malformations, and Dysmorphic Facial Features. Am J Hum Genet 104: 1073-1087.
Argyros O., Karampelas T., Varela A., Asvos X., Papakyriakou A., Agalou A., Beis D., Davos C. H., Fokas D., Tamvakopoulos C. (2017). Targeting of the breast cancer microenvironment with a potent and linkable oxindole based antiangiogenic small molecule. Oncotarget 8: 37250-37262.
Asimaki A., Kapoor S., Plovie E., Karin Arndt A., Adams E., Liu Z.Z., James C. A., Judge D. P., Calkins H., Churko J., Wu J. C., MacRae C. A., Kléber A. G., Saffitz J. E. (2014). Identification of a New Modulator of the Intercalated Disc in a Zebrafish Model of Arrhythmogenic Cardiomyopathy. Sci. Transl. Med. 6: 240ra74.
Beis D., Bartman T., Jin S.W., Scott I. C., D'Amico L. A., Ober E. A., Verkade H., Frantsve J., Field H. A., Wehman A., Baier H., Tallafuss A., Bally-Cuif L., Chen J.N., Stainier D. Y. R., Jungblut B. (2005). Genetic and cellular analyses of zebrafish atrioventricular cushion and valve development. Development 132: 4193-4204.
Beis D., Stainier D. Y.R. (2006). In vivo cell biology: following the zebrafish trend. Trends Cell Biol. 16: 105-112.
Bensimon-Brito A., Ramkumar S., Boezio G. L.M., Guenther S., Kuenne C., Helker C. S.M., Sánchez-Iranzo H., Iloska D., Piesker J., Pullamsetti S., Mercader N., Beis D., Stainier D. Y.R. (2020). TGF-β Signaling Promotes Tissue Formation during Cardiac Valve Regeneration in Adult Zebrafish. Dev Cell. 52: 9-20.e7.
Bobori D., Dimitriadi A., Karasiali S., Tsoumaki-Tsouroufli P., Mastora M., Kastrinaki G., Feidantsis K., Printzi A., Koumoundouros G., Kaloyianni M. (2020). Common mechanisms activated in the tissues of aquatic and terrestrial animal models after TiO2 nanoparticles exposure. Environ Int. 138: 105611.
Bournele D., Beis D. (2016). Zebrafish models of cardiovascular disease. Heart Fail Rev 21: 803-813.
Chaita E., Lambrinidis G., Cheimonidi C., Agalou A., Beis D., Trougakos I., Mikros E., Skaltsounis A.L., Aligiannis N. (2017). Anti-Melanogenic Properties of Greek Plants. A Novel Depigmenting Agent from Morus alba Wood. Molecules 22: 514.
G. Chatzimitakos T., Pliatsika C., Chousidis I., D. Leonardos I., Stalikas C. D. (2021). Metabolomic Profiling Unveils the Impact of Non-Doped and Heteroatom-Doped Carbon Nanodots on Zebrafish (Danio rerio) Embryos. Nanomaterials 11: 483.
Chousidis I., Chatzimitakos T., Leonardos D., Filiou M. D., Stalikas C. D., Leonardos I. D. (2020). Cannabinol in the spotlight: Toxicometabolomic study and behavioral analysis of zebrafish embryos exposed to the unknown cannabinoid. Chemosphere 252: 126417.
Chousidis I., Stalikas C. D., Leonardos I. D. (2020). Induced toxicity in early-life stage zebrafish (Danio rerio) and its behavioral analysis after exposure to non-doped, nitrogen-doped and nitrogen, sulfur-co doped carbon quantum dots. Environ Toxicol Pharmacol 79: 103426.
Christou M., Iliopoulou M., Witten P. E., Koumoundouros G. (2018). Segmentation pattern of zebrafish caudal fin is affected by developmental temperature and defined by multiple fusions between segments. J Exp Zool (Mol Dev Evol) 330: 330-340.
Costa B., Estrada M. F., Mendes R. V., Fior R. (2020). Zebrafish Avatars towards Personalized Medicine—A Comparative Review between Avatar Models. Cells 9: 293.
Damalas D. E., Bletsou A. A., Agalou A., Beis D., Thomaidis N. S. (2018). Assessment of the Acute Toxicity, Uptake and Biotransformation Potential of Benzotriazoles in Zebrafish ( Danio rerio ) Larvae Combining HILIC- with RPLC-HRMS for High-Throughput Identification . Environ. Sci. Technol. 52: 6023-6031.
Diamantis D. A., Agalou A., Chatziathanasiadou M. V., Markopoulos G. S., Bellou S., Kanaki Z., Crook T., Syed N., Rampias T., Klinakis A., Kolettas E., Beis D., Tzakos A. G. (2021). Biotin-Yellow a biotin guided NIR turn-on fluorescent probe for cancer targeted diagnosis. Sens Actuators B Chem. 337: 129807.
Dimitriadi A., Beis D., Arvanitidis C., Adriaens D., Koumoundouros G. (2018). Developmental temperature has persistent, sexually dimorphic effects on zebrafish cardiac anatomy. Sci Rep 8: 8125.
Dimitriadi A., Geladakis G., Koumoundouros G. (2021). 3D heart morphological changes in response to developmental temperature in zebrafish: More than ventricle roundness . J Morphol 282: 80-87.
Emmanouilidou A., Karetsou Z., Tzima E., Kobayashi T., Papamarcaki T. (2013). Knockdown of prothymosin α leads to apoptosis and developmental defects in zebrafish embryos. Biochem. Cell Biol. 91: 325-332.
Farrell J. A., Wang Y., Riesenfeld S. J., Shekhar K., Regev A., Schier A. F. (2018). Single-cell reconstruction of developmental trajectories during zebrafish embryogenesis. Science 360: eaar3131.
Fior R., Póvoa V., Mendes R. V., Carvalho T., Gomes A., Figueiredo N., Ferreira M. G. (2017). Single-cell functional and chemosensitive profiling of combinatorial colorectal therapy in zebrafish xenografts. Proc Natl Acad Sci USA 114: E8234-E8243.
Georga I., Koumoundouros G. (2010). Thermally induced plasticity of body shape in adult zebrafish Danio rerio (Hamilton, 1822). J. Morphol. 271: 1319-1327.
Giardoglou P., Beis D. (2019). On Zebrafish Disease Models and Matters of the Heart. Biomedicines 7: 15.
Giardoglou P., Bournele D., Park M., Kanoni S., Dedoussis G. V., Steinberg S. F., Deloukas P., Beis D. (2021). A zebrafish forward genetic screen identifies an indispensable threonine residue in the kinase domain of PRKD2. Biol Open 10: bio058542.
Huisken J., Swoger J., Del Bene F., Wittbrodt J., Stelzer E. H. K. (2004). Optical Sectioning Deep Inside Live Embryos by Selective Plane Illumination Microscopy. Science 305: 1007-1009.
Kakouri E., Agalou A., Kanakis C., Beis D., Tarantilis P. A. (2020). Crocins from Crocus sativus L. in the Management of Hyperglycemia. In Vivo Evidence from Zebrafish. Molecules 25: 5223.
Kalogirou S., Malissovas N., Moro E., Argenton F., Stainier D. Y.R., Beis D. (2014). Intracardiac flow dynamics regulate atrioventricular valve morphogenesis. Cardiovasc Res 104: 49-60.
Kaloyianni M., Dimitriadi A., Ovezik M., Stamkopoulou D., Feidantsis K., Kastrinaki G., Gallios G., Tsiaoussis I., Koumoundouros G., Bobori D. (2020). Magnetite nanoparticles effects on adverse responses of aquatic and terrestrial animal models. J Hazard Mater. 383: 121204.
Katharios P., Kalatzis P. G., Kokkari C., Pavlidis M., Wang Q. (2019). Characterization of a Highly Virulent Edwardsiella anguillarum Strain Isolated From Greek Aquaculture, and a Spontaneously Induced Prophage Therein. Front. Microbiol. 10: 141.
Katraki-Pavlou S., Kastana P., Bousis D., Ntenekou D., Varela A., Davos C. H., Nikou S., Papadaki E., Tsigkas G., Athanasiadis E., Herradon G., Mikelis C. M., Beis D., Papadimitriou E., (2022). Protein tyrosine phosphatase receptor-ζ1 deletion triggers defective heart morphogenesis in mice and zebrafish. American Journal of Physiology-Heart and Circulatory Physiology 322: H8-H24.
Katsouda A., Peleli M., Asimakopoulou A., Papapetropoulos A., Beis D., (2020). Generation and Characterization of a CRISPR/Cas9—Induced 3-mst Deficient Zebrafish. Biomolecules 10: 317.
Kefalos P., Agalou A., Kawakami K., Beis D. (2019). Reactivation of Notch signaling is required for cardiac valve regeneration. Sci Rep 9: 16059.
Kellaris G., Khan K., Baig S. M., Tsai I.C., Zamora F. M., Ruggieri P., Natowicz M. R., Katsanis N. (2018). A hypomorphic inherited pathogenic variant in DDX3X causes male intellectual disability with additional neurodevelopmental and neurodegenerative features. Hum Genomics 12: 11.
Keller P. J., Schmidt A. D., Wittbrodt J., Stelzer E. H.K. (2008). Reconstruction of Zebrafish Early Embryonic Development by Scanned Light Sheet Microscopy. Science 322: 1065-1069.
Lazari D., Alexiou G. A., Markopoulos G. S., Vartholomatos E., Hodaj E., Chousidis I., Leonardos I., Galani V., Kyritsis A. P. (2017). N-(p-coumaroyl) serotonin inhibits glioblastoma cells growth through triggering S-phase arrest and apoptosis. J Neurooncol 132: 373-381.
Malissovas N., Griffin L. B., Antonellis A., Beis D. (2016). Dimerization is required for GARS-mediated neurotoxicity in dominant CMT disease. Hum. Mol. Genet. 25: 1528-1542.
Moro E., Ozhan-Kizil G., Mongera A., Beis D., Wierzbicki C., Young R. M., Bournele D., Domenichini A., Valdivia L. E., Lum L., Chen C., Amatruda J. F., Tiso N., Weidinger G., Argenton F. (2012). In vivo Wnt signaling tracing through a transgenic biosensor fish reveals novel activity domains. Dev Biol 366: 327-340.
- OECD (2013). Test No. 236: Fish Embryo Acute Toxicity (FET) Test, OECD Guidelines for the Testing of Chemicals, Section 2. Home Page. https://www.oecd-ilibrary.org/environment/test-no-236-fish-embryo-acute-toxicity-fet-test_9789264203709-en
Orfanidou T., Papaefthimiou C., Antonopoulou E., Leonardos I., Theophilidis G. (2013). The force of the spontaneously contracting zebrafish heart, in the assessment of cardiovascular toxicity: Application on adriamycin. Toxicol In Vitro 27: 1440-1444.
Papadimitriou T., Armeni E., Stalikas C. D., Kagalou I., Leonardos I. D. (2012). Detection of microcystins in Pamvotis lake water and assessment of cyanobacterial bloom toxicity. Environ Monit Assess 184: 3043-3052.
Papakyriakou A., Kefalos P., Sarantis P., Tsiamantas C., Xanthopoulos K. P., Vourloumis D., Beis D. (2014). A Zebrafish In Vivo Phenotypic Assay to Identify 3-Aminothiophene-2-Carboxylic Acid-Based Angiogenesis Inhibitors . Assay Drug Dev Technol 12: 527-535.
Pavlidis M., Digka N., Theodoridi A., Campo A., Barsakis K., Skouradakis G., Samaras A., Tsalafouta A. (2013). Husbandry of Zebrafish, Danio Rerio , and the Cortisol Stress Response . Zebrafish 10: 524-531.
Pavlidis M., Theodoridi A., Tsalafouta A. (2015). Neuroendocrine regulation of the stress response in adult zebrafish, Danio rerio. Prog Neuropsychopharmacol Biol Psychiatry 60: 121-131.
Printzi A., Fragkoulis S., Dimitriadi A., Keklikoglou K., Arvanitidis C., Witten P. E., Koumoundouros G. (2021). Exercise‐induced lordosis in zebrafish Danio rerio (Hamilton, 1822) . J Fish Biol 98: 987-994.
Protonotarios N., Tsatsopoulou A., Patsourakos P., Alexopoulos D., Gezerlis P., Simitsis S., Scampardonis G. (1986). Cardiac abnormalities in familial palmoplantar keratosis.. Heart 56: 321-326.
Pyriochou A., Beis D., Koika V., Potytarchou C., Papadimitriou E., Zhou Z., Papapetropoulos A. (2006). Soluble Guanylyl Cyclase Activation Promotes Angiogenesis. J Pharmacol Exp Ther 319: 663-671.
Renieri E. A., Sfakianakis D. G., Alegakis A. A., Safenkova I. V., Buha A., Matović V., Tzardi M., Dzantiev B. B., Divanach P., Kentouri M., Tsatsakis A. M. (2017). Nonlinear responses to waterborne cadmium exposure in zebrafish. An in vivo study. Environ Res 157: 173-181.
Samaras A., Pavlidis M. (2020). A Modified Protocol for Cortisol Analysis in Zebrafish ( Danio rerio ), Individual Embryos, and Larvae . Zebrafish 17: 394-399.
Sarantis P., Gaitanaki C., Beis D. (2019). Ventricular remodeling of single-chambered myh6−/− adult zebrafish hearts occurs via a hyperplastic response and is accompanied by elastin deposition in the atrium. Cell Tissue Res 378: 279-288.
Serafimidis I., Heximer S., Beis D., Gavalas A. (2011). G Protein-Coupled Receptor Signaling and Sphingosine-1-Phosphate Play a Phylogenetically Conserved Role in Endocrine Pancreas Morphogenesis. Mol Cell Biol 31: 4442-4453.
Serifi I., Tzima E., Bardouki H., Lampri E., Papamarcaki T. (2019). Effects of the Essential Oil from Pistacia lentiscus Var. chia on the Lateral Line System and the Gene Expression Profile of Zebrafish (Danio rerio). Molecules 24: 3919.
Serifi I., Tzima E., Soupsana K., Karetsou Z., Beis D., Papamarcaki T. (2016). The zebrafish homologs of SET/I2PP2A oncoprotein: expression patterns and insights into their physiological roles during development. Biochem J. 473: 4609-4627.
Tsata V., Möllmert S., Schweitzer C., Kolb J., Möckel C., Böhm B., Rosso G., Lange C., Lesche M., Hammer J., Kesavan G., Beis D., Guck J., Brand M., Wehner D. (2021). A switch in pdgfrb cell-derived ECM composition prevents inhibitory scarring and promotes axon regeneration in the zebrafish spinal cord. Dev Cell. 56: 509-524.e9.
Tzima E., Serifi I., Tsikari I., Alzualde A., Leonardos I., Papamarcaki T. (2017). Transcriptional and Behavioral Responses of Zebrafish Larvae to Microcystin-LR Exposure. Int J Mol Sci 18: 365.
Vartholomatos E., Alexiou G. A., Markopoulos G. S., Lazari D., Tsiftsoglou O., Chousidis I., Leonardos I., Kyritsis A. P. (2020). Deglucohellebrin: A Potent Agent for Glioblastoma Treatment. Anticancer Agents Med Chem 20: 103-110.
Wagner D. E., Weinreb C., Collins Z. M., Briggs J. A., Megason S. G., Klein A. M. (2018). Single-cell mapping of gene expression landscapes and lineage in the zebrafish embryo. Science 360: 981-987.
Zoupa M., Machera K. (2017). Zebrafish as an Alternative Vertebrate Model for Investigating Developmental Toxicity—The Triadimefon Example. Int J Mol Sci 18: 817.
Zoupa M., Zwart E. P., Gremmer E. R., Nugraha A., Compeer S., Slob W., van der Ven L. T.M. (2020). Dose addition in chemical mixtures inducing craniofacial malformations in zebrafish (Danio rerio) embryos. Food and Chemical Toxicology 137: 111117.